library(maptools)
library(gstat)
library(geoR)
library(rgdal)
library(lattice)
library(RSAGA)
# MGI Balkans zone 6:
gk_6 <- "+proj=tmerc +lat_0=0 +lon_0=18 +k=0.9999 +x_0=6500000 +y_0=0 +ellps=bessel
+units=m +towgs84=550.499,164.116,475.142,5.80967,2.07902,-11.62386,0.99999445824"
# ------------------------------------------------------------
# Import of data to R:
# ------------------------------------------------------------
download.file("http://www.spatial-analyst.net/DATA/elevations.zip",
destfile=paste(getwd(), "elevations.zip", sep="/"))
for(j in list(".shp", ".shx", ".dbf")){
fname <- zip.file.extract(file=paste("elevations", j, sep=""), zipname="elevations.zip")
file.copy(fname, paste("./elevations", j, sep=""), overwrite=TRUE)
}
unlink("elevations.zip")
list.files(getwd(), recursive=T, full=F)
# import the map to R:
elevations <- readShapePoints("elevations.shp", proj4string=CRS(gk_6))
str(elevations)
names(elevations@data) <- "Z"
# ------------------------------------------------------------
# Variogram modelling:
# ------------------------------------------------------------
range(elevations$Z)
# sub-sample - geoR can not deal with large datasets!
sel <- runif(length(elevations@data[[1]]))<0.2
Z.geo <- as.geodata(elevations[sel,"Z"])
# histogram:
plot(Z.geo, qt.col=grey(runif(4)))
# Variogram modelling (target variable):
par(mfrow=c(1,2))
# anisotropy:
plot(variog4(Z.geo, max.dist=1000, messages=FALSE), lwd=2)
# fit variogram using likfit:
Z.svar <- variog(Z.geo, max.dist=1000, messages=FALSE)
Z.vgm <- variofit(Z.svar, messages=FALSE, ini=c(var(Z.geo$data), 1000), fix.nugget=T, nugget=0)
Z.vgm
# confidence bands:
env.model <- variog.model.env(Z.geo, obj.var=Z.svar, model=Z.vgm)
plot(Z.svar, envelope=env.model); lines(Z.vgm, lwd=2);
dev.off()
# spacing between contours:
bin.Z <- (max(elevations$Z)-min(elevations$Z))*(length(elevations$Z))^(-1/3)
round(bin.Z, 0)
# ------------------------------------------------------------
# Geostatistical simulations:
# ------------------------------------------------------------
# prepare an empty grid:
demgrid <- spsample(elevations, type="regular", cellsize=c(30,30))
gridded(demgrid) <- TRUE
fullgrid(demgrid) <- TRUE
demgrid@grid
gridcell <- demgrid@grid@cellsize[1]
# locs <- pred_grid(c(demgrid@bbox[1,1]+gridcell/2, demgrid@bbox[1,2]-gridcell/2),
c(demgrid@bbox[2,1]+gridcell/2, demgrid@bbox[2,2]-gridcell/2), by=gridcell)
# conditional simulations:
# Z.sims <- krige.conv(Z.geo, locations=locs, krige=krige.control(obj.m=Z.vgm), output=output.control(n.predictive=1))
# this is computationally very intensive; geoR is simply not fit to work with large data!
# copy the values fitted in geoR:
Z.ovgm <- vgm(psill=Z.vgm$cov.pars[1], model="Mat", range=Z.vgm$cov.pars[2], nugget=Z.vgm$nugget, kappa=1.2)
# *Kappa is artificially set at 1.2! to produce smooth surface!
Z.ovgm
# conditional simulations in gstat:
N.sim <- 100
DEM.sim <- krige(Z~1, elevations, demgrid, Z.ovgm, nmax=30, nsim=N.sim)
fullgrid(DEM.sim) <- TRUE
# plot 4 realizations:
spplot(DEM.sim[1:4], col.regions=grey(seq(0,1,0.025)))
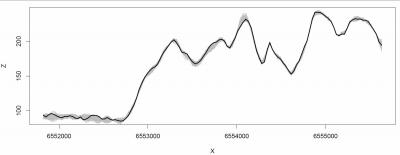
# Cross-section at y=5,073,012:
cross.s <- data.frame(X=seq(demgrid@bbox[1,1]+gridcell/2, demgrid@bbox[1,2]-gridcell/2, gridcell),
Y=rep(5073012, demgrid@[email protected][1]))
coordinates(cross.s) <-~X+Y
# proj4string(cross.s) <- elevations@proj4string
cross.ov <- overlay(DEM.sim, cross.s)
plot(cross.ov@coords[,1], cross.ov@data[[1]], type="l", xlab="X", ylab="Z", col="grey")
for(i in 2:N.sim-1){
lines(cross.ov@coords[,1], cross.ov@data[[i]], col="grey")
}
lines(cross.ov@coords[,1], cross.ov@data[[N.sim]], lwd=2)


