Who's onlineThere are currently 0 users and 4 guests online.
User loginBook navigationNavigationLive Traffic MapNew Publications
|
Fig. 7.5: Soil Organic Carbon stock (kg C m^2) for South America.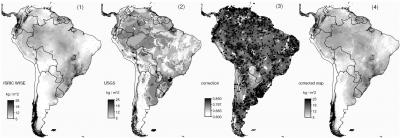
library(maptools)
library(rgdal)
library(RSAGA)
library(gstat)
SA.aea <- "+proj=aea +lat_1=-5 +lat_2=-42 +lat_0=-32 +lon_0=-60 +x_0=0 +y_0=0 +ellps=aust_SA +units=m +no_defs"
# load the gridded and point data:
load(url("http://spatial-analyst.net/book/system/files/WISE_SOC.Rdata"))
SA.bbox <- gridsSA@bbox
writeOGR(GSPD.aea, "GSPD_aea.shp", ".", "ESRI Shapefile")
# subset the points:
rsaga.geoprocessor(lib="shapes_tools", module=14, param=list(SHAPES="GSPD_aea.shp",
CUT="m_GSPD_aea.shp", METHOD=0, TARGET=0, CUT_AX=SA.bbox[1,1],
CUT_BX=SA.bbox[1,2], CUT_AY=SA.bbox[2,1], CUT_BY=SA.bbox[2,2]))
# ------------------------------------------------------------
# Variogram modelling
# ------------------------------------------------------------
# original variable:
m_SOC <- readShapePoints("m_GSPD_aea.shp", CRS(SA.aea))
m_SOC.ov <- overlay(gridsSA, m_SOC)
m_SOC.ov$SOC <- m_SOC$SOC
m_SOC.ov <- remove.duplicates(m_SOC.ov) # there are duplicate points!
sel <- !is.na(m_SOC.ov$SOC)&!is.na(m_SOC.ov$SOC_reg)
SOC.svar <- variogram(log1p(SOC)~1, m_SOC.ov[sel,])
SOC.ovgm <- fit.variogram(SOC.svar, vgm(nugget=0, model="Exp", range=80000, sill=var(log1p(m_SOC.ov$SOC), na.rm=T)))
SOC.ovgm.plt <- plot(SOC.svar, SOC.ovgm, pc="+", cex=1.5, pl=TRUE, col="black", main="Original variable")
# residuals:
res_SOC.svar <- variogram(log1p(SOC)~SOC_reg, m_SOC.ov[sel,])
SOC.rvgm <- fit.variogram(res_SOC.svar, vgm(nugget=var(m_SOC.ov$SOC, na.rm=T)/2,
model="Exp", range=80000, sill=var(m_SOC.ov$SOC, na.rm=T)/2))
SOC.rvgm.plt <- plot(res_SOC.svar, SOC.rvgm, pc="+", cex=1.5, pl=FALSE, col="black", main="Residuals")
SOC.rvgm
# synchronize the two plots:
SOC.rvgm.plt$x.limits <- SOC.ovgm.plt$x.limits
SOC.rvgm.plt$y.limits <- SOC.ovgm.plt$y.limits
print(SOC.ovgm.plt, split=c(1,1,2,1), more=TRUE)
print(SOC.rvgm.plt, split=c(2,1,2,1), more=FALSE)
# regression-kriging:
m_SOC.rk <- krige(log1p(SOC)~SOC_reg, m_SOC.ov[sel,], gridsSA, SOC.rvgm, nmin=30, nmax=40, block=c(10e3, 10e3))
# back-transform:
m_SOC.rk$SOC_rk <- expm1(m_SOC.rk$var1.pred)
# Correct the map where the density of points is low using the USDA map:
download.file("http://spatial-analyst.net/worldmaps/SOC.zip", destfile=paste(getwd(), "/SOC.zip", sep=""))
unzip("SOC.zip")
# resample to the same grid:
rsaga.esri.to.sgrd(in.grids="SOC.asc", out.sgrd="SOC.sgrd", in.path=getwd())
rsaga.geoprocessor(lib="pj_proj4", 2, param=list(SOURCE_PROJ="\"+proj=longlat +datum=WGS84\"",
TARGET_PROJ=paste('"', SA.aea ,'"', sep=""), SOURCE="SOC.sgrd", TARGET="m_SOC_USDA.sgrd", TARGET_TYPE=2,
INTERPOLATION=1, GET_SYSTEM_SYSTEM_NX=m_SOC.rk@[email protected][[1]], GET_SYSTEM_SYSTEM_NY=m_SOC.rk@[email protected][[2]],
GET_SYSTEM_SYSTEM_X=m_SOC.rk@[email protected][[1]], GET_SYSTEM_SYSTEM_Y=m_SOC.rk@[email protected][[2]],
GET_SYSTEM_SYSTEM_D=10000))
rsaga.sgrd.to.esri(in.sgrds="m_SOC_USDA.sgrd", out.grids="m_SOC_USDA.asc", out.path=getwd(), prec=3)
m_SOC.rk$SOC_USDA <- readGDAL("m_SOC_USDA.asc")$band1
# merge the two maps (BCSP formula):
w <- sqrt(m_SOC.rk$var1.var)/sqrt(var(log1p(m_SOC.ov$SOC), na.rm=T))
m_SOC.rk$w <- 1-w/max(w, na.rm=TRUE)
m_SOC.rk$SOC.f <- round((m_SOC.rk$w * m_SOC.rk$SOC_rk+(1-m_SOC.rk$w) * m_SOC.rk$SOC_USDA), 0)
spplot(m_SOC.rk[c("SOC_rk","SOC_USDA", "SOC.f")], col.regions=bpy.colors(), at=seq(0,60,5))
|
Testimonials"From a period in which geographic information systems, and later geocomputation and geographical information science, have been agenda setters, there seems to be interest in trying things out, in expressing ideas in code, and in encouraging others to apply the coded functions in teaching and applied research settings." Poll |




Recent comments
8 years 47 weeks ago
9 years 12 weeks ago
9 years 21 weeks ago
9 years 33 weeks ago